Transformed Gaussian Markov Random Field
tgmrf.RdUse this function to fit spatial data using non-guassian copulas.
tgmrf(data, formula, neigh, scale = T, spatial_var, neigh_order = NULL, group_var = NULL, change_vars = NULL, beta = NULL, nu = 1, eps = NULL, mu = NULL, rho_s = 0, rho_t = 0, rho_st = 0, tau = NULL, family = "poisson", type = "gamma-shape", mat_type = "car", method = "metropolis", nsim = 1000, burnin = 0, thin = 1, E = NULL, n = 1, prior_param = NULL, MCMC_config = NULL, fix_rho = NULL, range = list(rho_s = c(-1, 1), rho_t = c(-1, 1), rho_st = c(-1, 1)), verbose = FALSE, c_beta = NULL, c_eps = NULL, c_mu = NULL, c_nu = NULL, c_rho = NULL, conj_beta = TRUE)
Arguments
| data | A data.frame with all variables used to fit a model |
|---|---|
| formula | Formula to fit the model |
| neigh | Neighborhood structure of nb class or a neighborhood matrix |
| scale | Logical parameter to scale covariates |
| spatial_var | Variable name that indicates the column which represents the spatial variable |
| neigh_order | Order that the spatial_var appear on data |
| group_var | Variable name that indicates the column which represents the variable variable |
| change_vars | Variables that change in the time. We'll return n_var parameters for this variable. One for each n_var. |
| beta | Vector of the initial values to coeficients vector |
| nu | A Initial value to variance of copula |
| rho_s | Spatial dependence parameter |
| rho_t | Temporal dependence parameter |
| rho_st | Spatio-temporal dependence parameter |
| family | 'Poisson' or 'Binary' |
| type | Depends of family. 'lognormal', 'lognormal-precision', 'gamma-shape', 'gamma-scale', 'weibull-shape', 'weibull-scale' for Poisson family 'beta-logit', 'beta-probit', 'beta-alpha', 'beta-beta' for Binomial family |
| mat_type | car or leroux |
| method | metropolis or arms |
| nsim | Number of MCMC iterations |
| burnin | number of discards iterations |
| thin | Lag to collect the observations |
| E | Offsets for Poisson data |
| n | Number of trials in Binomial data |
| prior_param | List of priors parameters. Gaussian for beta vector and Gamma for nu. Entrys are: 'nu' (list with shape and scale) and 'beta' (list with mean and precision) |
| MCMC_config | Parameters to tunning MCMC. Entrys are: 'arms' (list with ninit and maxpoint) and 'metropolis' (list with var_beta, var_eps, var_rho and var_log_nu) |
| fix_rho | A list informing if any rho (rho_s, rho_t, rho_st) is fixed |
| range | A list informing a range for sampling rho (rho_s, rho_t, rho_st). Each entre must be a vector in the interval (-1, 1). |
Value
beta A matrix with samples of beta
nu A vector with samples of nu
rho A vector with samples of rho
Examples
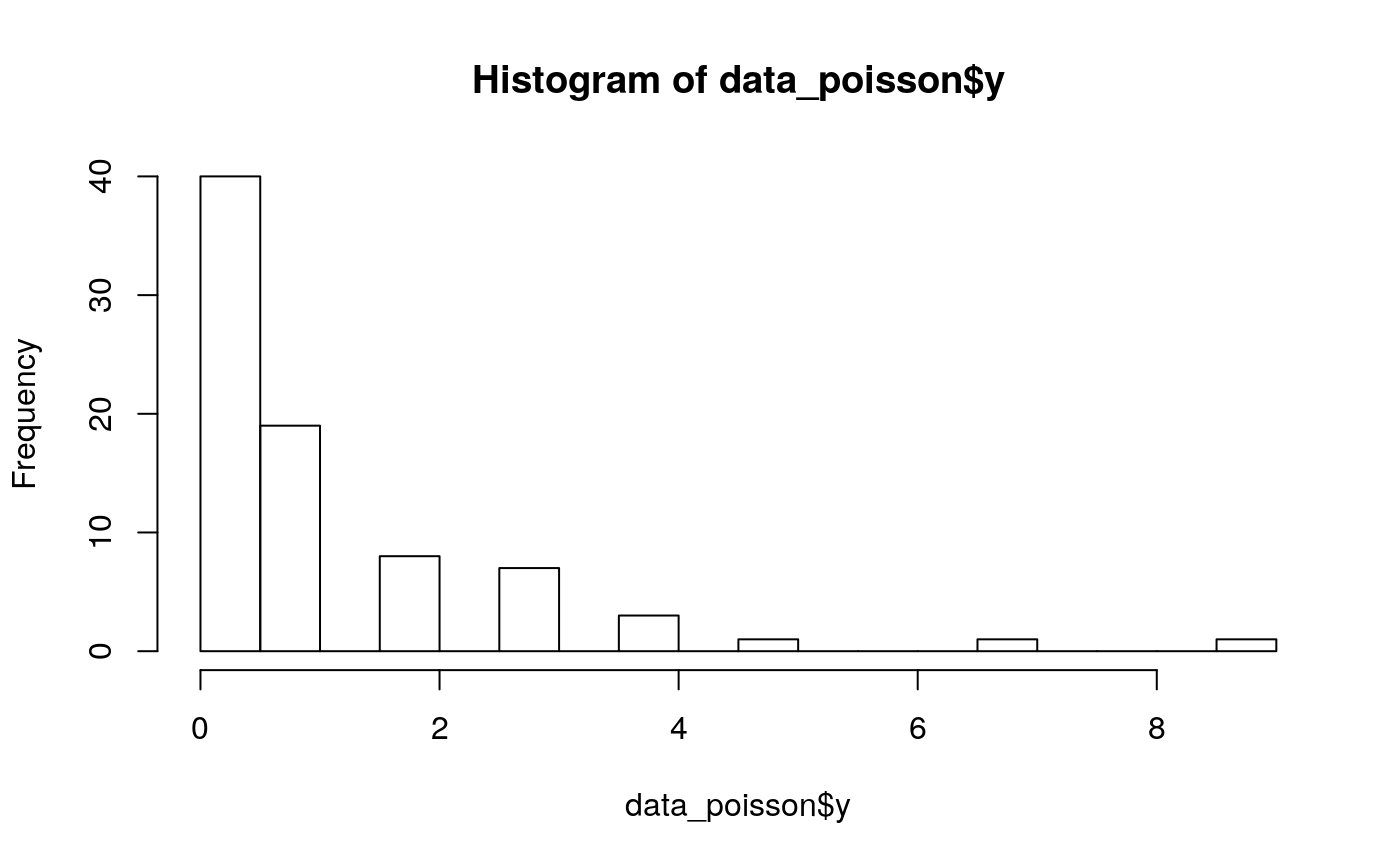
#>#>#>#>#>library(truncnorm) library(OpenMPController) #'#'-- Configurações ---- rowid <- 8 colid <- 5 n_vars <- 2 N <- rowid*colid*n_vars betas <- 0.5 intercept <- -.5 nu <- 0.3 P <- length(betas) X <- scale(matrix(c(rnorm(n = rowid*colid*n_vars, mean = 0)), ncol = P)) type_data <- 'lognormal' family <- 'poisson' seed <- 123456 rho_s <- 0.95*2.25 rho_t <- 0.00001 rho_st <- 0.00001 #'#'-- Dados ---- data_poisson <- rtgmrf(rowid = rowid, colid = colid, X = X, n_var = n_vars, rho_s = rho_s, rho_t = rho_t, rho_st = rho_st, betas = betas, nu = nu, intercept = intercept, type_data = type_data, family = family, seed = seed) hist(data_poisson$y, breaks = 20)bd <- data.frame(y = data_poisson$y, regiao = data_poisson$reg, grupo = data_poisson$var, X1 = data_poisson$X, eps = data_poisson$eps, check.names = F) neigh <- data_poisson$neigh #'#'-- Configurações ---- nsim <- 20000 burnin <- 0 thin <- 1 formula <- paste0('y ~ X1') #'#'-- Metropolis rcpp ---- vars_beta <- diag(c(0.03, 0.02), nrow = P+1, ncol = P+1) vars_eps <- diag(0.0001, nrow = N, ncol = N) var_log_nu <- 0.85 var_rho <- diag(c(0.1, 0.3, 0.05), ncol = 3, nrow = 3) omp_set_num_threads(n = 1)#> [[1]] #> [1] 1 #>type_model <- type_data c_beta <- 1#'(2.38^2)/P c_eps <- 1#'(2.38^2)/N c_nu <- 1#'(2.38^2) c_rho <- 1#'(2.38^2)/3 system.time( out_metcpp <- tgmrf(data = bd, formula = formula, spatial_var = "regiao", group_var = "grupo", beta = c(intercept, betas), eps = data_poisson$eps, nu = nu, rho_s = 0.95, rho_t = 0, rho_st = 0, family = family, type = type_model, mat_type = "car", method = "metropolis", nsim = nsim, burnin = burnin, thin = thin, prior_param = list("nu" = list("shape" = 0.2, "rate" = 0.2)), E = NULL, neigh = neigh, fix_rho = list("rho_s" = FALSE, "rho_t" = FALSE, "rho_st" = FALSE), MCMC_config = list('metropolis' = list('var_beta' = vars_beta, 'var_eps' = vars_eps, 'var_log_nu' = var_log_nu, 'var_rho' = var_rho)), range = list("rho_s" = c(-1, 1), "rho_t" = c(-1, 1), "rho_st" = c(-1, 1)), scale = FALSE, verbose = TRUE, c_beta = c_beta, c_eps = c_eps, c_nu = c_nu, c_rho = c_rho) )#> Error in poimcar_cpp(nsim = nsim, burnin = burnin, thin = thin, eps = eps, mu = mu, beta = beta, nu = nu, rho_s = rho_s, rho_t = rho_t, rho_st = rho_st, X = X, y = y, E = E, Ws = Ws, Wt = Wt, N = N, P = P, mean_beta = mean_beta, tau_beta = tau_beta, eta_nu = eta_nu, psi_nu = psi_nu, fix_rho_s = fix_rho_s, fix_rho_t = fix_rho_t, fix_rho_st = fix_rho_st, range_rho_s = range_rho_s, range_rho_t = range_rho_t, range_rho_st = range_rho_st, type = type_num, var_beta_met = var_beta, var_eps_met = var_eps, var_log_mu_met = var_log_mu, var_log_nu_met = var_log_nu, var_rho_met = var_rho, verbose = verbose, c_beta = c_beta, c_eps = c_eps, c_mu = c_mu, c_nu = c_nu, c_rho = c_rho, conj_beta = conj_beta): Mat::operator(): index out of bounds#>summary(out_metcpp)#> Error in summary(out_metcpp): object 'out_metcpp' not found